Highlight
Microfluidic Device for High Throughput RNA Splice Variation Studies
Achievement/Results
Nick Carroll, a Trainee supported by the NSF IGERT program on Integrating Nanotechnology with Cell Biology and Neuroscience (INCBN), is working at the University of New Mexico on development of a technology to quantitatively measure the relative abundance of all splice variants of a specific gene in a cancer sample. The objective is to measure the exon expression of all exons in the context of a complete structure of the spliced mRNA. The work focuses on development of a complete microfluidic device, or “Lab-on-a-Chip” system, for the high-throughput analysis of splice variant patterns in hundreds of samples per day. The Lab-on-a-Chip system will automate the measurements so that quantitative and contextual information about alternative RNA splicing can be obtained from large numbers of patient samples.
Microfluidic devices have the advantage of allowing generation of monodispersed emulsions with droplet sizes from ~10 µm up to nearly 1 mm in diameter. For efficiently loading large amplicons onto beads, a large emulsion droplet size is a very imporant capability. Additionally, microfluidic devices are highly valuable for studying clinical samples, where quantities are limited. The group of Prof. Dimiter Petsev, Nick Carroll’s main advisor, can generate emulsions with very small amounts of starting material, on the order of a few microliters (lower limit is less than 1 µl), whereas the bulk emulsions require hundreds of microliters (at same concentration as the microfluidic samples – thus hundreds of times higher amounts of starting material). Finally, the labor is greatly reduced for running a microfluidic device relative to using bulk emulsions, due to potential automation. Ultimately, the microfluidic device will be interfaced to an autosampler to load multiple samples sequentially.
In addition to optimizing droplet size, bead loading in the microfluidic droplets has been maximized for DNA amplification and immobilization. Proper droplet loading (with beads) is highly nontrivial. Under the direction of his INCBN IGERT co-mentor, Prof. Jeremy Edwards, Nick Carroll has been able to successfully overcome many of these difficulties, and currently can properly load between 40 and 50% of all droplets with beads. Large DNA molecule loading on the beads b has also been optimized by using different types of beads.
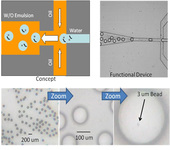
Fig. 1 shows the microfluidic device for generating monodispersed emulsion with large drop size (~100 µm). This size is not possible using bulk emulsion methods (i.e. homogenizer). The bottom row shows the size of the monodispersed emulsion drops. The lower right image shows that it is possible to create and load the drops in a microfluidic device. In this experiment, the drops were generated at a rate of 100/s.
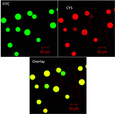
Following amplification and immobilization of complimentary DNA molecules onto polysaccharide agarose beads, fluorescent probes were hybridized to certain regions of the single stranded DNA to test for the existence (or absence) of certain exons. A particular emission wavelength detected corresponds to a specific exon type. Thus, detection of this wavelength can be used to verify the existence of a specific exon contained within the amplified cDNA, as illustrated in Figure 2. The green fluorescence (FITC) probe is a control to verify the ligation of the cDNA to the bead via a primer, while the red (Cy5) fluorescence indicates the presence of exon 9 in the c-Myb gene. The bottom image is the overlay of these colors.
Address Goals
This activity primarily fosters research that will advance the frontiers of knowledge, establishing the Nation as a global leader in high throughput, inexpensive DNA sequencing. The ability to selectively introduce and control chemical reagents, biomolecules, and cells within microfluidic devices facilitates a new generation of experiments and assays. The benefits of using microfluidic devices for laboratory applications include:
- Reduction in sample volume and reagent usage
- Improved resolution of separations
- Ability to run reaction and analysis processes faster
- Ability to run processes in parallel
- Improved control of mixing and heating of fluids
- Rapid mass transfer as a result of high surface area to volume ratios
- Improved integration of process steps, for example reactions and separations
- Development of new and improved detection methods
- Simpler and cheaper disposable devices
- Access to a wide range of fluidic geometries